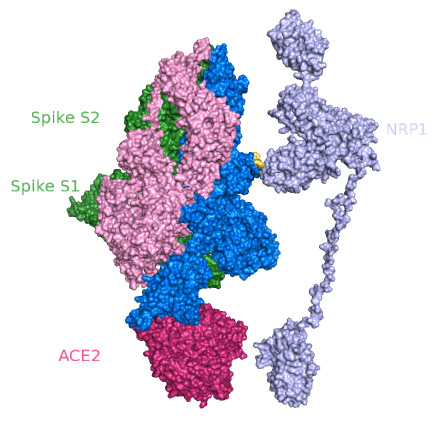
Despite being co-translationally inserted in the ER membrane, most M proteins do not contain a signal sequence. It has a small N-terminal glycosylated ectodomain and a much larger C-terminal endodomain that extends 6–8 nm into the viral particle. It is a small (~25–30 kDa) protein with three transmembrane domains and is thought to give the virion its shape. The M protein is the most abundant structural protein in the virion. S1 makes up the large receptor-binding domain of the S protein, while S2 forms the stalk of the spike molecule. In most, coronaviruses, S is cleaved by a host cell furin-like protease into two separate polypeptides noted S1 and S2. The trimeric S glycoprotein is a class I fusion protein and mediates attachment to the host receptor. Homotrimers of the virus encoded S protein make up the distinctive spike structure on the surface of the virus. The S protein (~150 kDa), utilizes an N-terminal signal sequence to gain access to the ER, and is heavily N-linked glycosylated. These are the spike (S), membrane (M), envelope (E), and nucleocapsid (N) proteins, all of which are encoded within the 3′ end of the viral genome. Coronaviruses have helically symmetrical nucleocapsids, which is uncommon among positive-sense RNA viruses, but far more common for negative-sense RNA viruses.Ĭoronavirus particles contain four main structural proteins. Within the envelope of the virion is the nucleocapsid. These spikes are a defining feature of the virion and give them the appearance of a solar corona, prompting the name, coronaviruses. The most prominent feature of coronaviruses is the club-shaped spike projections emanating from the surface of the virion. Ĭoronavirus virions are spherical with diameters of approximately 125 nm as depicted in recent studies by cryo-electron tomography and cryo-electron microscopy. The accessory proteins are almost exclusively nonessential for replication in tissue culture however, some have been shown to have important roles in viral pathogenesis. The organization of the coronavirus genome is 5′-leader-UTR-replicase-S (Spike)-E (Envelope)-M (Membrane)-N (Nucleocapsid)-3′ UTR-poly (A) tail with accessory genes interspersed within the structural genes at the 3′ end of the genome (Fig. The 3′ UTR also contains RNA structures required for replication and synthesis of viral RNA. Additionally, at the beginning of each structural or accessory gene are transcriptional regulatory sequences (TRSs) that are required for expression of each of these genes ( see Subheading 4.3 on RNA replication). The 5′ end of the genome contains a leader sequence and untranslated region (UTR) that contains multiple stem loop structures required for RNA replication and transcription. The replicase gene encoding the non-structural proteins (nsps) occupies two-thirds of the genome, about 20 kb, as opposed to the structural and accessory proteins, which make up only about 10 kb of the viral genome. The genome contains a 5′ cap structure along with a 3′ poly (A) tail, allowing it to act as an mRNA for translation of the replicase polyproteins. These differences cause significant alterations in the structure and morphology of the nucleocapsids and virions.Ĭoronaviruses contain a non-segmented, positive-sense RNA genome of ~30 kb. In fact, the Nidovirales order name is derived from these nested 3′ mRNAs as nido is Latin for “nest.” The major differences within the Nidovirus families are in the number, type, and sizes of the structural proteins.  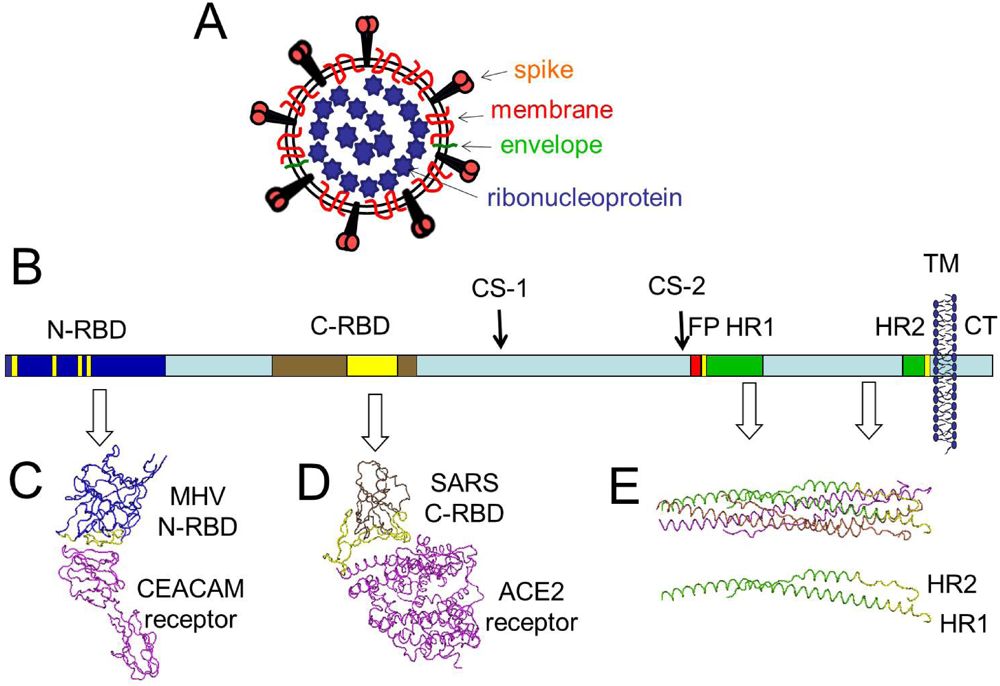
Other common features within the Nidovirales order include: (1) a highly conserved genomic organization, with a large replicase gene preceding structural and accessory genes (2) expression of many non-structural genes by ribosomal frameshifting (3) several unique or unusual enzymatic activities encoded within the large replicase–transcriptase polyprotein and (4) expression of downstream genes by synthesis of 3′ nested sub-genomic mRNAs. They all contain very large genomes for RNA viruses, with some viruses having the largest identified RNA genomes, containing up to 33.5 kilobase (kb) genomes. The viruses were initially sorted into these genera based on serology but are now divided by phylogenetic clustering.Īll viruses in the Nidovirales order are enveloped, non-segmented positive-sense RNA viruses. 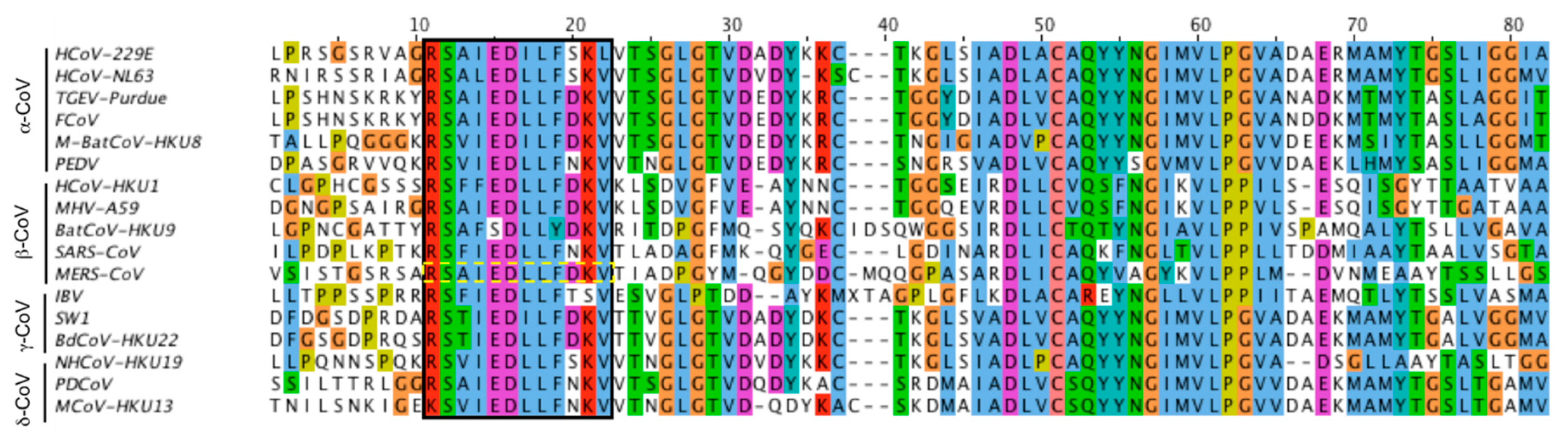
The Coronavirinae are further subdivided into four genera, the alpha, beta, gamma, and delta coronaviruses. The Coronavirinae comprise one of two subfamilies in the Coronaviridae family, with the other being the Torovirinae. Coronaviruses (CoVs) are the largest group of viruses belonging to the Nidovirales order, which includes Coronaviridae, Arteriviridae, Mesoniviridae, and Roniviridae families.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
